Hormones and Behavior
, September 2020
Conditional inactivation of Npy1r gene in mice induces sex-related differences of metabolic and behavioral function
Ilaria Bertocchi abc1 , Alessandra Oberto abc1 , Angela Longo a , Paola Palanza d , Carola Eva abc
Sex hormone-driven differences in gene expression have been identified in experimental animals, highlighting brain neuronal populations implicated in dimorphism of metabolic and behavioral functions. Neuropeptide Y-Y1 receptor (NPY-Y1R) system is sexually dimorphic and sensitive to gonadal steroids. In the present study we compared the phenotype of male and female conditional knockout mice ( Npy1r rfb mice), carrying the inactivation of Npy1r gene in excitatory neurons of the brain limbic system.
Compared to their male control (Npy1r 2lox ) littermates, male Npy1r rfb mice exhibited hyperactivation of the hypothalamic–pituitary–adrenal (HPA) axis that is associated with anxiety and executive dysfunction, reduced body weight growth, after-fasting refeeding, white adipose tissue (WAT) mass and plasma leptin levels. Conversely, female Npy1r rfb mice displayed an anxious-like behavior but no differences in HPA axis activity, executive function and body weight, compared to control females.
Moreover, conditional inactivation of Npy1r gene induced an increase of subcutaneous and gonadal WAT weight and plasma leptin levels and a compensatory decrease of Agouti-related protein immunoreactivity in the hypothalamic arcuate (ARC) nucleus in females, compared to their respective control littermates. Interestingly, Npy1r mRNA expression was reduced in the ARC and in the paraventricular hypothalamic nuclei of female, but not male mice.
These results demonstrated that female mice are resilient to hormonal and metabolic effects of limbic Npy1r gene inactivation, suggesting the existence of an estrogen-dependent relay necessary to ensure the maintenance of the homeostasis, that can be mediated by hypothalamic Y1R.
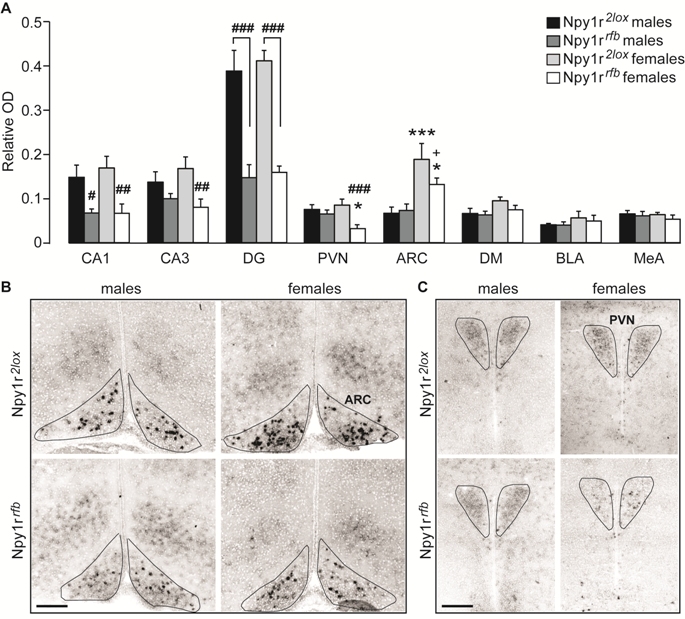
Fig. 1. Expression of Npy1r mRNA in the brain of Npy1r 2lox and Npy1r rfb mice of both sexes. (A) Semiquantitative in situ hybridization signal intensity (relative OD) analysis revealed a significant decreaseof Npy1r mRNA expression inCA1, CA3 and DG cell bodies of maleand female Npy1r rfb mice and in thePVN and ARC of female but not of maleNpy1r rfb mice, compared with their sexpaired control littermates. A significantincrease of Npy1r mRNA was observedin the ARC of female mice comparedwith their male counterparts, independentlyof the genotype. CA1, CA2and CA3: ###p < 0.001,##p < 0.01 and #p < 0.05 versussex paired Npy1r 2lox mice, by Tukeytest. PVN: ###p = 0.001 versus femaleNpy1r 2lox mice and *p < 0.05versus male Npy1r rfb mice by Tukey.ARC: ***p < 0.001 versus maleNpy1r 2lox mice; *p < 0.05 versusmale Npy1r rfb mice and +p = 0.051versus female Npy1r 2lox mice by Tukeytest. Data are the mean ± SEM.n = 8–6 from 4 to 7 litters. (B and C)Representative picture showing Npy1rmRNA signal intensity differences inthe ARC (B) and in the PVN (C) of maleand female Npy1r 2lox and Npy1r rfb mice. The black lines were drawn onthe digitized images to trace theboundaries selected nuclei (based onthe cellular density differences withthe surrounding structures) and to definethe area of interest (AOI) used for quantitation of mRNA expression. Scale bars: 250 μm. CA1, cornu ammonis 1 hippocampal subregion; CA3, cornu ammonis3 hippocampal subregion; DG, dentate gyrus; PVN, paraventricular nucleus of hypothalamus; ARC, arcuate nucleus of hypothalamus; DM, dorsomedial nucleus ofhypothalamus; BLA, basolateral nucleus of amygdala; MeA, medial nucleus of amygdala.
a
Neuroscience Institute of the Cavalieri-Ottolenghi Foundation, 10043 Orbassano, Turin, Italy
b
Department of Neuroscience, University of Turin, 10126 Turin, Italy
c
Neuroscience Institute of Turin, Italy
d
Department of Medicine and Surgery, University of Parma, 43100 Parma, Italy